Serologic Antibody Assays
Initial testing for the diagnosis of hepatitis C infection uses serologic assays that detect human antibodies generated as a response to hepatitis C virus (HCV) infection.[1,2,3] A positive HCV antibody test indicates one of the following three scenarios: (1) active infection, (2) past HCV infection that has resolved or been cured, or (3) a false-positive test.[1,4] None of these anti-HCV antibody tests can differentiate whether the infection is new (acute), chronic, or no longer present.
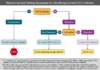
- Enzyme Immunoassay (EIA): The third-generation HCV EIA detects antibodies that bind to recombinant antigens derived from four HCV regions: core, nonstructural 3, nonstructural 4, and nonstructural 5 (Figure 1).[6,7] The EIA test is reported as positive or negative based on an absorbance signal compared with a cutoff value.
- Sensitivity of EIA and False-Negative Results: The third-generation HCV EIA has a sensitivity of approximately 98%.[8,9,10] Circumstances associated with a false-negative EIA include patients with acute HCV infection, persons with major immune compromising conditions (advanced HIV infection or organ transplantation recipients), and persons with chronic renal failure on long-term hemodialysis.
- Specificity of EIA and False-Positive Results: The third-generation HCV EIA has a reported specificity greater than 99%; false-positive tests can occur with increased gamma globulin production, with autoimmune diseases, and following immunizations.[8] In addition, a false-positive test is more likely when performing widespread testing in populations that have a very low HCV prevalence.
- Chemiluminescence Immunoassay (CIA): The CIA test is an antibody test similar to the EIA but is used less frequently than the EIA test. For the diagnosis of HCV, the CIA has similar sensitivity and specificity as the third-generation EIA.[1,11]
- Point-of-Care Rapid Immunoassays: The OraQuick HCV Rapid Antibody Test was approved by the U.S. Food and Drug Administration (FDA) in 2010 as a point-of-care test for use with whole blood samples obtained by either venipuncture or fingerstick. This OraQuick Rapid Antibody Test can be used as an alternative to the third-generation EIA for initial HCV antibody testing.[12,13,14,15] The OraQuick test is read 20 to 40 minutes after the test device is inserted into the buffer, and the result is either reactive or nonreactive (Figure 2).[12,13] In 2011, the FDA granted a Clinical Laboratory Improvement Amendments (CLIA) waiver for the OraQuick HCV Rapid Antibody Test. Additional point-of-care rapid HCV antibody tests have been developed but are not approved for use in the United States.[16,17]
Molecular HCV RNA Tests
Molecular diagnostic tests for hepatitis C specifically detect HCV RNA and the process is commonly referred to as a Nucleic Acid Test (NAT) or Nucleic Acid Amplification Test (NAAT).[18] The HCV NAT becomes positive approximately 1 to 2 weeks after initial HCV infection.[19] The NAT test has become the gold standard supplemental test for patients who have a positive HCV EIA screening test.[2,18] The NAT can determine whether a patient with a positive HCV antibody test has current (active) or resolved HCV infection.[2,6] In addition, the NAT can be used in combination with other laboratory studies, such as prior antibody test results or hepatic aminotransferase levels, to suggest the possibility of acute HCV infection.[19] The results for the commercially available quantitative HCV RNA assays, which were previously reported as copies/mL, are now given in International Units per milliliter (IUs/mL).[6]
- Qualitative HCV RNA: The qualitative HCV RNA tests provide a yes or no answer to whether detectable HCV RNA is present in the patient's blood sample.[20] The qualitative HCV RNA assays are FDA-approved for HCV diagnostic purposes. These tests, however, do not provide a quantitative level of HCV and are not used for baseline HCV RNA levels or for monitoring response to therapy.[20] For most qualitative HCV RNA assays, the lower limit of detection is 10-15 IU/mL.[20]
- Quantitative HCV RNA: The quantitative HCV RNA tests detect and quantify the number of HCV copies in the patient's blood sample, reported as IU/mL. Clinically, these tests are used for diagnosing HCV and monitoring response to therapy. Quantitative HCV RNA tests used for diagnosis and monitoring should have a lower limit of detection of 25 IU/mL or less.[6,21]
Immunoassays for HCV Core Antigen
As an HCV diagnostic marker, HCV core antigen has been studied, either alone or as an HCV antibody-HCV antigen combination assay.[22,23] Some experts have proposed the use of an HCV core antigen test as a less expensive option than HCV RNA testing, but there are no HCV antigen assays (or HCV antigen-antibody combination assays) that are FDA-approved for use in the United States at this time.[24,25]
 Elbasvir-Grazoprevir Zepatier
Elbasvir-Grazoprevir Zepatier Glecaprevir-Pibrentasvir Mavyret
Glecaprevir-Pibrentasvir Mavyret Ledipasvir-Sofosbuvir Harvoni
Ledipasvir-Sofosbuvir Harvoni Ribavirin Copegus, Rebetol, Ribasphere
Ribavirin Copegus, Rebetol, Ribasphere Sofosbuvir Sovaldi
Sofosbuvir Sovaldi Sofosbuvir-Velpatasvir Epclusa
Sofosbuvir-Velpatasvir Epclusa Sofosbuvir-Velpatasvir-Voxilaprevir Vosevi
Sofosbuvir-Velpatasvir-Voxilaprevir Vosevi