genome architecture analysis
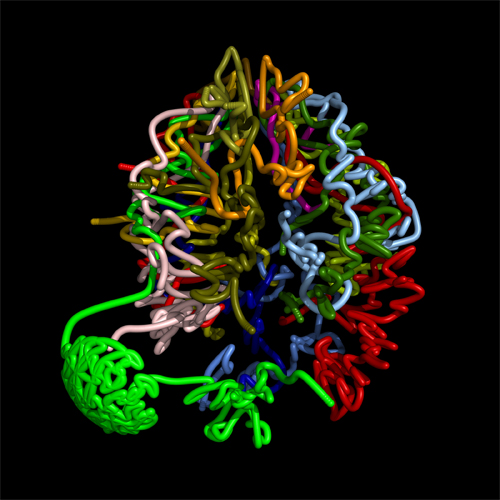
One of the fundamental questions of the post-genome era is: How does genotype (an organism's genetic structure) translate into phenotype (an observable characteristic)? Our genetic information is contained in 23 pairs of chromosomes. The 46 chromosomes are arranged not in straight lines, but in a tangle, like a plate of cooked spaghetti. The result is that a gene's nearest neighbors in three-dimensional space may be on completely different chromosomes. In collaboration with the laboratories of Dr. Bill Noble, Dr. Stan Fields, and Dr. Jay Shendure (all faculty members in the Department of Genome Sciences), Zhijun Duan in our lab developed a method for capturing interactions within and between chromosomes genome-wide. Mirela Andronescu (Noble Lab) used this information to construct a three-dimensional model of the genome of budding yeast (Duan et al., Nature)
The information provides an entirely new epigenetic view of the genome, and will likely provide important insights into cell state, cell identity, cancer, and evolution. We plan to use this new method to characterize the genomic architecture of stem cells, and to understand how this changes during stem cell differentiation.
A Three-Dimensional Model of the Yeast Genome

Paper
Duan Z, Andronescu M, Schutz K, McIlwain S, Kim YJ, Lee C, Shendure J, Fields S, Blau CA*, Noble WS*. A three dimensional model of the yeast genome. Nature 2010 May 20;465(7296):363-7. Epub 2010 May 2. (* denotes co-corresponding authors)