DeepTracer, Data Analysis and Machine Learning for 3D Electron Microscopy
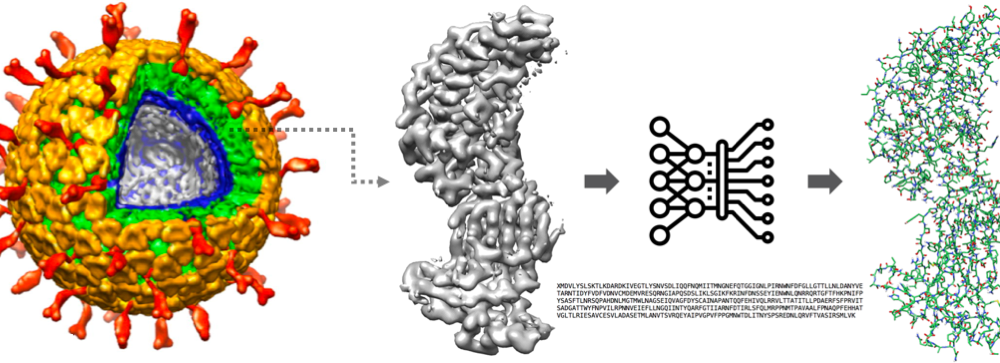
We combine software development (front- and back-end), 3D image processing, machine learning, data mining, and geometric modeling techniques for automatic and accurate protein structure prediction based on cutting-edge new technology – Electron cryo-microscopy (cryo-EM).
Background: Life ultimately depends on the interactions of large biological molecules, such as viruses. The nature of these interactions depends on the 3D shape and structure of these molecules. Cryo-EM as a cutting-edge technology has carved a niche for itself in the study of large-scale protein complexes. However, it is still challenging to detect the protein structures automatically and accurately from the 3D EM volume data.
Project Goals/Outcomes:
- Prediction tools and software for the Bio-medicine community;
- Smart frameworks for mining large-scale 3D volume data;
- Interactive and user-friendly platform for structure modeling and data visualization
Student Outcomes: Collaborative teamwork, programming skills, problem-solving skills, publication experiences, etc.
Requirements:
- Proficient programming and software development skills (Python, GitHub, etc.)
- Foundations in Data Structures, Algorithms, and OOP
- Good understanding of 3D geometry
- Passion for interdisciplinary research and learning new concepts
- Understand, review, and survey the existing literature in 3D visual data analysis and machine learning
- Collaborate with other group members on the testing and implementation of prediction and data analysis algorithms
Time commitment:
- Minimum commitment of 5 hours a week for 2 quarters with the registration of CSS497, CSS499, or other independent study or faculty research credits.
- Attend weekly research group meetings.
Schools or Related Disciplines:
Business
Educational Studies
First Year and Pre-major Program (FYPP)
Interdisciplinary Arts and Sciences (IAS)
Nursing and Health Studies (NHS)
Science, Technology, Engineering and Math (STEM)
STEM – Computing and Software Systems (CSS)
STEM – Engineering and Mathematics
Category: Research and Creative Projects
Time: estimated hours per week is 1hr – 9hrs
Best for student levels (to apply and/or participate): High school, Undergraduate students, post-Grad, Graduate level, or Alumni
Credit/Compensation Notes: Academic credit available. This is a volunteer or unpaid position. Sometimes hourly pay or awards/stipends are available.
Contact: Dong Si, Ph.D., dongsi@uw.edu
Go to project or opportunity website for more information
