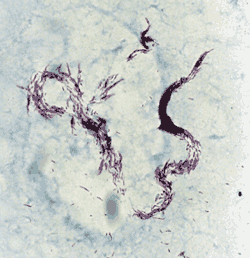
Accurate identification of mycobacterial isolates is an essential task of the clinical microbiology laboratory that enables initiation of proper antimicrobial therapy. For many organisms, traditional phenotypic identification may be difficult, laborious and time-consuming. This issue is confounded by phenotypic variation within species, the many newly described pathogenic species and the limited battery of phenotypic tests available to distinguish among all established and potential bacterial pathogens. It is sometimes essential to identify bacterial isolates to species level in order to detect unsuspected pathogens, ascribe pathogenicity to species so far considered to be nonpathogenic and to identify new bacterial species.
With >500000 sequences available in public databases, 16S rRNA gene sequencing is considered by current taxonomists to be the gold standard in identification and classification of bacteria, including the slower growing mycobacteria. It contains conserved regions useful for the design of broad-range primers that amplify the gene from all pathogenic and nonpathogenic bacteria in addition to hypervariable regions that contain species-specific signature sequences useful for bacterial identification to species level.
For rapidly growing mycobacteria (RGM), the 65-KDa heat shock protein and RNA polymerase Beta subunit genes are more variable than the 16S rRNA gene, making these targets useful in the discrimiation of closely related species such as M. abscessus, M. chelonae, M. bolletii, and M. massiliense. Among the RGM, there are important differences in antimicrobial susceptibilities and virulence. Hsp65 and rpoB gene sequencing generates direct, unambiguous data and can distinguish medically relevant subspecific phylogenetic lineages. The UWMC Molecular Diagnosis Sequence Database has been validated with complete phenotypic characterization, including morphological, biochemical, HPLC and GLC for fatty acid analysis, as well as susceptibility testing.