On Friday, August 9th, the Bernier Lab participated in Seattle’s 2nd Rare Disease Fair. The National Organization for Rare Diseases (NORD) member organization, BORN A HERO, hosted this rare disease awareness event in collaboration with Seattle Children’s Research Institute. They invited individuals to come learn about what makes a disease or disorder “rare,” learn about different rare medical conditions, what questions remain unanswered, and what we can do about it. It was such a great event and we are proud to be part of this amazing network of parents, advocates, researchers and medical professionals. We look forward to continued collaborations and helping rare disease!
On Friday, August 9th, the Bernier Lab participated in Seattle’s 2nd Rare Disease Fair. The National Organization for Rare Diseases (NORD) member organization, BORN A HERO, hosted this rare disease awareness event in collaboration with Seattle Children’s Research Institute. They invited individuals to come learn about what makes a disease or disorder “rare,” learn about different rare medical conditions, what questions remain unanswered, and what we can do about it. It was such a great event and we are proud to be part of this amazing network of parents, advocates, researchers and medical professionals. We look forward to continued collaborations and helping rare disease!

Category Archives: Uncategorized
Meet Bernier Lab’s Amazing Summer Intern, Claire
Hello everyone! My name is Claire Shaver. I am from Telluride, Colorado, located in the bottom south-west corner of CO and about a 6 hour drive from Denver. This upcoming school year I will be a senior in high school. Whenever not in school, you often see me playing as goalie on my high school soccer team or volunteering for a nonprofit in Telluride called Telluride Adaptive Sports Program. This summer I had the chance to be an intern at the Bernier Lab for 7 weeks due to an amazing Telluride-based program called Pinhead Institute. Pinhead Institute has an internship program, which strives to get high school students into STEM labs over the summer between their junior and senior year. I have had the privileged to represent this program this summer at the Bernier Lab.
I found and became interested in the Bernier Lab by the ABC-CT study they just recently complete. I was interested in the lab because they are able to study those with autism by interacting with them rather than studying different genetic components in a regular lab setting. Over the past 7 weeks, I have been able to see this interaction first hand by being able to observe different testing they complete in order to assess common phenotype (what you see) features of autism. I was also able to attend two family conferences they hosted, DYRK1A and SCN2A, in which I was able to interact with the participants, learn more about these genetic events, and learn more about how the Bernier Lab completes the testing.
Want to learn more about my time in Seattle at the Bernier Lab? Read my weekly blog posts in which I talk about my daily tasks in the lab and other fun-activities I did during my stay in Seattle! I want to thank all those who made this internship possible: to Sarah Holbrooke at Pinhead for coordinating my internship, to Micah and Eva at the lab for being my mentors and teaching me about the different aspects in the lab, and to my Seattle host family for allowing me to stay their house for these 7 weeks.
Local Seattle Attractions Cater to Those With Sensory Sensitivities
When diagnosed with autism, children often also receive a diagnosis of having a sensory sensitivity, meaning that they want to reduce or avoid loud noises, bright lights, and other over simulating events. When presented with over-stimulus, some children with ASD may shut down, become alarmed or agitated, or have other outward or inward behavior to express their discomfort. Given that the rates that children diagnosed with autism are rising, Seattle’s Woodland Park Zoo and Pacific Science Center wanted to create spaces that are designed for children with sensory sensitivity so they can enjoy these classic Seattle attractions.
The Woodland Park Zoo has created a Woodland Park Zoo Sensory Map which points out areas for the children to escape the crowds if needed (aka quiet areas), areas where they can run or move, and other attractions that cater to those with sensory sensitivities. Along with the sensory map, the zoo has also built a Sensory Garden in which children can play with different textures and different instruments.
Once a month the Pacific Science Center has a special early opening or late closing for those with sensory sensitivities in order to provide a calmer experience not only for the children but also for their caregivers. When sites provide accessibility to those living with disabilities, it opens the door to many opportunities that these families might have otherwise avoided. We are grateful to the Woodland Zoo and Pacific Science Center for taking initiative in making these attractions more accessibility for children with diverse needs.
Local Komo News covers how these classic Seattle attractions have adapted to accommodate children with special needs. Watch the video here.
Emotional problems and social success in ASD
In Emily Neuhaus and Sara Jane Webb’s article on Spectrum News titled, “For autistic children, emotional problems may hinder social success,” they discuss commonalities found in parent completed behavioral checklists about the relationship between social motivation and emotional challenges.
After looking at more than 2,000 behavioral checklists, Neuhaus and Webb found that two out of three children diagnosed with autism have difficulty to meet threshold for clinical evaluation in a certain area. Once comparing their scores with the parents’ reports, they were able to conclude that higher social motivation allows for better social success. They were also able to conclude that there is correlation between emotional problems (anxiety, irritability, or aggression) and poor social skills. Overall they concluded that children to be socially successful need to have social motivation and good emotional regulation. Based on these findings, they recommend that children with an ASD diagnosis and mild emotional difficulties would benefit from therapy in order to gather a set of tools for regulating their emotions during social interaction.
Top Autism Gene and Defining Characteristics
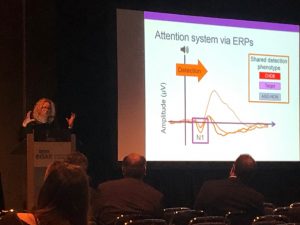
Last week our very own Dr. Caitlin Hudac presented unpublished data at the 2019 International Society for Autism Research (INSAR) annual meeting that showed how disruptive mutations either to CHD8 – a top autism gene – or genes CHD8 controls may define a subtype of Autism. These mutations have been seen to related to similar behavioral characteristics, like enlarged heads (Brenna Boyd in a poster at INSAR) and severe social problems (Dr. Jen Beighley and Dr. Hudac, under review). By looking at these mutations as a subtype, this will help to bridge how the impact of these genes at a molecular level therefore creates behavior that we can observe.
Through looking at children with disruptive CHD8 mutations or a mutation to one of six genes that CHD8 regulates, it was found that these mutations largely affects social behavior, everyday functioning/intelligence and unusually large head size. Compared to mutations of target genes, CHD8 participants tended to have the most social problems on average, but also had the best adaptive skills (Beighley, Hudac). Though timing of development of large head differs between CHD8 mutations and target genes (Boyd), they had common electroencephalography (EEG) brain responses which may reflect an ASD-associated early enhanced hypersensitivity but a lack of sustained response to certain noises. These findings could underlie some of the sensory sensitivities that characterize CHD8 and related genetic groups.
Full article can be found here on spectrum news: https://www.spectrumnews.org/news/targets-top-autism-gene-may-define-form-condition/
INSAR 2019
INSAR 2019 has begun, and the Bernier Lab showed to this years International Meeting for Autism Research, in Montreal, Canada, in amazing numbers! You can find our teams posters and talks on our website here at our webpage INSAR 2019. Here is a list of our team’s presentations:
At the International Society for Autism Research (INSAR) annual meeting, we will be presenting the following:
Thursday May 2, 2019
- Oral presentation: 2:05 pm in Room 516ABC
- Dr. Caitlin Hudac will be presenting a talk on “Characterizing the neural phenotype of CHD8 and CHD8-regulated targets”.
- Poster presentations: 5:30-7:30pm in Room 710
- Brenna Boyd will be presenting r on “A comparison of Head Circumference Growth Trajectories in the Context of the CHD8 Regulatory Network.”
- Monique Mahony will be presenting on “GroopIt: An Innovative Platform to Speed up Rare Genetic Disorder Research.”
- Sandy Trinh will be presenting on “Exploring Social Profiles of Individuals with 16p11.2 Deletion and Duplication.”
Friday May 3, 2019
- Poster presentations: 11:30 am – 1:30 pm
- Poster presentations: 5:30-7:30pm in Room 710
- Dr. Caitlin Hudac will be presenting on “Examining the Broader Autism Phenotype in the Context of Genetic Etiology.”
- Dr. Eva Kurtz-Nelson will be presenting on the “Clinical phenotype of de novo mutations in CHD2.”