Guarded Command Programming + Cell Signaling + Micro-Colony Simulation
What can you gro?
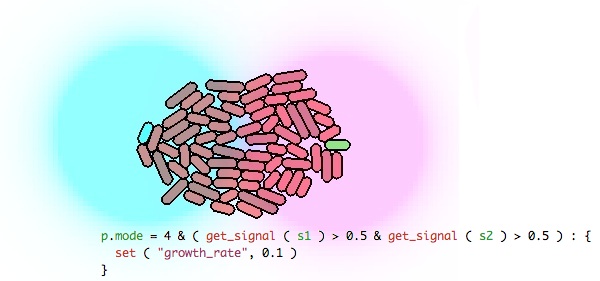 |
gro is a language for programming, modeling, specifying and simulating the behavior of cells in growing microcolonies of microorganisms. The simulator models cell growth, cell division, intrinsic and extrinsic noise, diffusing molecular signals, microchemostats, chemotaxis, and more. The gro framework is intended to be used in synthetic biology to prototype distributed, multicell behaviors and check that, logically, the local interaction rules you specify produce the desired global result. The language allows behaviors to be specified at whatever level of abstraction makes sense: from high level code, to low level biomolecular interations.
The gro framework has been also been used in the classroom, at UW and elsewhere, to teach synthetic biology to engineers. If you would like to learn more about gro, start by reading the documentation (see the link at the left). The tutorial, in particular, describes many of the main features of gro.
For examples, click on the Gallery link on the left, or visit our youtube channel!
Publications
M. Gutiérrez, P. Gregorio-Godoy, G. Pérez del Pulgar, L.E. Muñoz, S. Sáez, and A. Rodríguez-Patón, "A New Improved and Extended Version of the Multicell Bacterial Simulator gro" ACS Synthetic Biology, April, 2017.
K. Oishi and E. Klavins, "A Framework for Implementing Finite State Machines in Gene Regulatory Networks", ACS Synthetic Biology, Feb. 2014.
S.S. Jang, K.T. Oishi, R.G. Egbert, and E. Klavins, "Specification and simulation of multicelled behaviors", ACS Synthetic Biology, July, 2012.
Tutorial Slides
- Writing programs, modeling protein expression, working with data. (pptx), (pdf)
- Controlling the simulation, chemostats, program composition. (pptx), (pdf)
- Signals and reaction/diffusion equations. (pptx), (pdf)
- Saving frames and making movies. (pptx), (pdf)
 |
 |
Developed by The Klavins Lab,
University Washington, Seattle, WA
Copyright © University of Washington. All rights reserved.