As published in Dimensions, the Magazine of the UW Alzheimer's Disease Research Center - Fall 2017 (Download the PDF or Read Online)
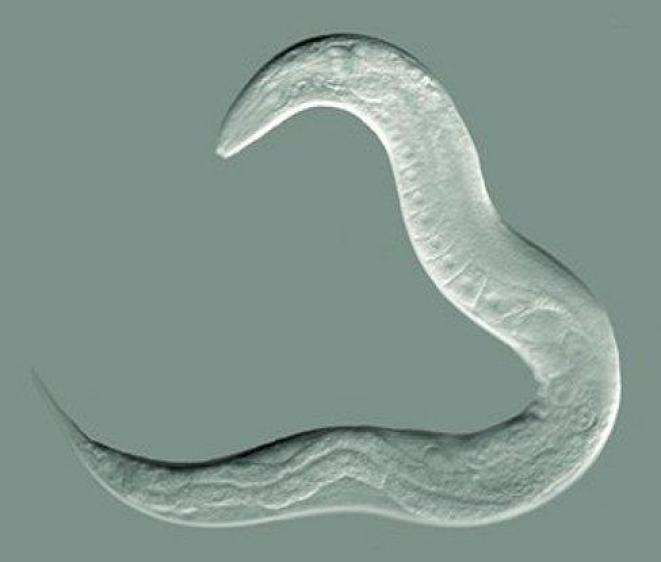
“Powerful.” “Fantastic.” “Cute.” “Rad.” “Far out.” This is how UW ADRC researchers have been overheard talking about the Caenorhabditis elegans roundworm. Why do they love this 1-mm creature?
First, scientists find these little worms to be a valuable model to study aspects of human diseases, such as Alzheimer’s. C. elegans has intestines, muscles, a pharynx, and a simple nervous system of 302 neurons, compared to our 100 billion. Because the body is transparent, researchers can use a light microscope to look at any tissue they want at a 200-nm resolution, which is enough to observe physiology in living detail.
To make a worm model of neurodegenerative disease, researchers insert a gene that triggers the worm’s genome to express one of the toxic proteins related to Alzheimer’s disease in humans—including tau, amyloid beta, TDP-43, or alpha synuclein. The toxicity of these proteins probably plays out similarly in C. elegans because much of the cellular machinery for processing proteins is similar across all animal species. The worms develop paralysis, delayed swallowing, and other ‘phenotypes’ of disease.
Researchers can use these worm models to quickly screen for any genes or compounds that modify the toxic effects of these pathologies or progression of diseases in the worm—a step towards finding therapeutic targets in humans.
Here is a tour through the different ways that worms play starring roles in UW Alzheimer's Disease Research Center (ADRC)-related projects in the UW Medicine’s Department of Pathology and Department of Geriatrics & Gerontology.
Exosomes are nano-sized packages, called vesicles, floating around many cell types, including neurons. They are like text messaging for cells, carrying genetic signals and proteins to distant areas of the body. They also scoop up waste in neurons, such as amyloid beta peptides, and clear it out. However, some scientists strongly suspect that some exosomes may spill some of their toxic cargo, contributing to the spreading of Alzheimer’s disease pathology in human patients.
The Kaeberlein Lab and collaborators are developing the simple roundworm, C. elegans, as a model to study how toxic Alzheimer’s disease-related proteins affect exosome function.
Through a collaborative effort with the groups led by Dr. Michael MacCoss in the UW Dept. of Genome Sciences, Dr. Dirk Keene in the UW Dept. of Pathology, and Dr. Alexander Mendenhall in the UW Dept. of Pathology, they uncovered that these worm vesicles act much like human exosomes. Their studies to date indicate that exosomes in Alzheimer’s disease-model worms also package up toxic proteins and send them out of neurons. This project was selected for a talk at the 21st International C. elegans meeting in June.
The Kaeberlein Lab’s Dr. Josh Russell and his collaborators in the ADRC Neuropathology Core use biotechnology to detect Alzheimer’s pathology in exosomes from worms and samples of cerebrospinal fluid from human patients. One laser-based tool, flow cytometry, suspends cells in a stream of fluid and passes them through an apparatus for counting or analysis. Ultimately, they want to find out if exosomes could be used in early diagnosis and treatment.
In healthy C. elegans, exosomes do something interesting. During the molting process, the worm’s skin cells bubble out exosomes, which then assemble into three stripes on each side of the worm’s body. Dr. Alex Mendenhall in the UW Dept. of Pathology says they look like “racing stripes” on a sports car.
In research stemming from an ADRC-funded project, the Mendenhall Lab team found that worms producing toxic proteins don’t form the racing stripes correctly. Other labs have observed this as well. Sometimes the stripes are absent, and sometimes they look strange—“like someone spilled the ink for the racing stripes,” says Mendenhall. This suggests that the Alzheimer’s-related proteins mess up exosomes’ ability to lay down the stripes.
This finding is exciting because they now have a phenotype of disrupted exosome function. They can screen for genes that have an effect on this phenotype. They will then knock out promising candidate genes in their worm models of Alzheimer’s disease, to see if symptoms get better or worse. This critical knowledge could point to ways to intervene in exosome biology to delay the progression of Alzheimer’s disease.
Think “multidisciplinary” is just an academic buzzword? ADRC researchers made a big discovery using bioinformatics, C. elegans worms, and postmortem human brains.
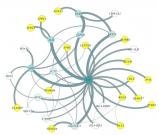
Dr. Shubhabrata Mukherjee, a statistical geneticist and Research Assistant Professor, UW Department of Medicine, used a systems biology computational approach to identify a network of genes involved in Alzheimer’s risk. The hub gene—or the gene with the highest degree of connections to other genes within the network—was UBC, a key gene in the ubiquitin pathway known to repair DNA.
Drs. Matt Kaeberlein and Josh Russell validated that UBC is linked to Alzheimer’s disease in their C. elegans worm models. And collaborators in the Ertekin-Taner lab at Mayo Clinic Jacksonville found reduced levels of UBC expression in those from people with Alzheimer’s disease. They conclude that UBC is an organizing member of this gene network, which may show decreased activity in people who eventually develop Alzheimer’s disease. The hope is that a therapeutic could target the ubiquitin pathway. Dr. Paul Crane was also involved in this work.
What initiates the early phases of neurodegeneration and its progression through brain cells? Research led by Dr. Brian Kraemer, Research Associate Professor in the UW Medicine Division of Gerontology, found new evidence that enzymes called kinases are integral to FTLD-TDP-43 disease progression in C. elegans worms.
Dr. Kraemer, along with Dr. Nicole Liachko, Acting Assistant Professor, and Laura Taylor, Graduate Research Associate, are working to understand why phosphorylation of TDP-43 goes wrong. “If we could ever prevent that modification, we would prevent these diseases from happening in the first place,” says Taylor.
Kraemer’s group has received new funding from the National Institute on Aging (NIA) for a project to study the role of the tau protein in Alzheimer’s disease, using a variety of model organisms of tau toxicity, including C. elegans worm, mouse, and human cells. Specifically, the grant allows the team to continue studying how the sut-2/MSUT2 gene controls tau aggregation and toxicity, and therefore determines a cell’s vulnerability to tau pathology. This knowledge may provide a novel candidate therapeutic target for pharmacological intervention.