BUILDING THE MOLECULAR GENETICS TEAM
With Bird ready to do the sleuthing necessary to find more families, and Schellenberg prepared to scour subjects' DNA for clues, they still needed someone to find the answers in the treasure trove of genetic information. At the prodding of George Martin, the Dean’s office stepped in to help and ultimately provided funding for the ADRC to recruit a young scientist from Stanford University, Ellen Wijsman, PhD. She was a statistical geneticist who was uniquely equipped to operate in the worlds of mathematics and computers, as well as in population and disease genetics. “Genius, that one,” notes Marie Walters. “I mean, I was in awe whenever I was with her.”
Wijsman’s job was crucial. She would use new computing technology to run genetic linkage analysis, combing through thousands of Schellenberg’s genotypes in search of strong correlations between the inheritance of DNA regions and the development of Alzheimer’s disease.
The UW ADRC’s pioneering project was on the right track, with the right team in place: Bird examined patients and diagnosed Alzheimer's, and ensured quality clinical data, with help from Ellen Nemens, who obtained blood samples and helped evaluate and maintain contact with families; Gerard Schellenberg, molecular biologist, handled the DNA work of generating genotypes; and statistical geneticist technician Ellen Wijsman developed computational statistical methods to analyze all the data collected from the research and search for genetic linkages to Alzheimer’s disease. Technicians in the ADRC’s Cell and Tissue Bank grew and maintained white blood cells—the source of DNA—from living patients' samples; and neuropathologist S. Mark Sumi, MD, the first leader of the Autopsy Core, along with others in UW Department of Pathology, studied the brain tissues from deceased participants.
Also connected to the effort were people such as Eric Larson, MD, MPH, Associate Professor of Medicine in the UW Department of Medicine, who ran the ADRC’s registry of patients living with familial and sporadic (non-familial) Alzheimer’s who consented for eventual brain autopsy, complemented by a data bank of the clinical information on the subjects. This resource enabled work on many ADRC research questions. For example, Murray Raskind, MD, Associate Professor in the UW Department of Psychiatry and Behavior Sciences, soon to be joined by Elaine Peskind, MD, studied whether neurochemicals found in cerebrospinal fluid samples were different between familial and sporadic Alzheimer’s patients.
Though the ADRC team were among the first in their fields to dive into Alzheimer’s from the genetic angle, they were not the only ones looking for an Alzheimer’s gene.
In 1985, researchers George Glenner, MD of University of San Diego, and Colin Masters, MD of University of Melbourne, isolated the plaques seen in the brains of people with Alzheimer’s disease and found that they were made of amyloid beta protein. “So, researchers started looking for the gene that coded for amyloid beta,” said Bird. “It turned out to be on chromosome 21, the same chromosome that is present in excess in Down's syndrome.” This finding explained a curious connection between Down’s syndrome and Alzheimer’s disease.
For years, scientists had known that virtually all individuals with Down’s syndrome eventually develop the hallmark plaques and tangles of Alzheimer’s disease by age 40, and some, but not all, later develop dementia. As word of the finding spread, other researchers looked at their families with Alzheimer’s disease and indeed came out with results showing a genetic linkage to chromosome 21.
The team at the UW ADRC followed suit in search of this linkage to chromosome 21 in their families. “We were looking for any joint inheritance of the disease with a single genetic marker, and we were using the developing gene mapping resources of the Human Genome Project,” says Wijsman. “Others were building maps and figuring out where these markers were. As they mapped those markers, our team would use them for linkage analysis to query each little region of the genome in search of a link to disease.”
But the science didn’t happen overnight. Bird set a high bar for enrolling patient families into the study, holding on to the same standard of requiring clear evidence that at least two patients per family had proven Alzheimer’s disease, based on autopsy and tissue analysis.
The genotyping was another tall order. In 1988, the technology allowed Schellenberg to genotype about 1,000 genetic markers in a week—but there were many medical pedigrees to work through. Covering the genome to the best of their ability meant that each individual needed genotyping for nearly 200 genetic markers each.
Running linkage analysis on each one of those markers, one at a time, for their entire study population, added up to a huge computational challenge. “Back when I was getting started at the ADRC, I had a single PC that was completely inadequate. It had one Central Processing Unit, and one processor. It was slow, and some of our analyses took six months,” says Wijsman, who is now Lead of the ADRC Data Management and Statistics Core. “I was borrowing computer time from anyone I could find that could spare it.”
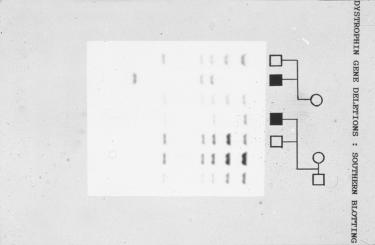
'Restriction fragment length polymorphism' technology (left) was used to genotype individuals represented on a medical pedigree (right). Courtesy, Wessex Reg. Genetics Centre
"It's a real team approach—people with very special expertise,” said Bird. “That's why it takes a lot of time and costs a lot of money."
The ADRC researchers rigorously studied pieces of DNA on chromosome 21 from the families in their cohort, including the Volga German family, but they did not find the linkage that other studies had found. "There are a couple of possible reasons why we haven't found the linkage," Bird said at the time in 1989. "The Alzheimer's disease in our families may be genetic, but it may be caused by a different gene than in other families. Or perhaps all genetic Alzheimer's is on chromosome 21, but there are presently too few genetic markers to look at, so we haven't been able to see the connection yet. We're just not sure."
London researchers would identify the elusive mutation in the APP gene, located on chromosome 21, in 1991 and showed that it was the cause of Alzheimer’s disease in two families in England and America. In the years leading up to this finding, the scientific community at large surmised that all cases of familial Alzheimer’s disease would be caused by this single gene on chromosome 21, but fortunately, some researchers at the UW ADRC were not so certain.
A SHARPER FOCUS
While many in the scientific community were rallying around the easy link between chromosome 21 and Alzheimer’s disease, Ellen Wijsman was ready with a healthy dose of skepticism. “As a population geneticist, you're aware of how much genetic diversity probably exists underneath the biological hood of one disease category,” says Wijsman. “When I started studying Alzheimer's and working with Tom and Jerry, I very quickly became convinced that there was going to be more than one gene for familial Alzheimer’s disease – that this was going to be more complicated than the field thought it would be.”
The slow pace of research ended up being a blessing in disguise—she had the time and bandwidth to think critically. “It took a long time to generate the genotype data, so I had a lot of time to think about our study design,” says Wijsman. She crunched the numbers and found that it would take years to run statistical tests and look at genotypes for one genetic marker at a time for each case. Additionally, Wijsman realized that if there were multiple different Alzheimer’s genes in their study population, then they would compete for attention in the linkage analysis; it would be difficult if not impossible to get a strong signal for one genetic disease factor. “I sat down and started assessing the drawbacks of trying to do everything all at once, versus recognizing that there may be more than one gene and being prepared for that possibility,” says Wijsman.

Ellen Wijsman, PhD in the early 1990s
Confronting these realities, Wijsman called a meeting and made her case. “It was very obvious to me that we really needed to change our study design,” she said. She made an argument to start dividing the patients up into groups based on how they differed in clinical presentation. Wijsman wanted to focus on the early-onset cases, based on the availability of detailed medical data about the very first signs of disease and nature of symptoms. She split the cases up according to an under-appreciated variable at the time— how old they were the last time their family reported that they were clinically normal."
As it turned out, focusing on age-of-onset was a useful way to differentiate between the disease’s familial forms. When they divided the patients up by age of symptom onset, they found that the late-onset families didn’t seem to follow any strong pattern of inheritance.
“I saw that the later-onset families didn’t always have a parent with Alzheimer’s and then a bunch of adult children with it as well,” says Wijsman. For the early-onset families, however, it looked like the offspring of a parent with Alzheimer’s had been born with a 50/50 chance of inheriting Alzheimer’s. The team reasoned that first focusing on the genetically straight-forward early-onset population would help them to interpret the data.
“While everybody else in the field was looking at all of their families at the same time, we started focusing in on these subpopulations and really using the data as efficiently as possible,” says Wijsman.
The team moved their research in a pioneering direction. In 1990, Bird published a paper that shared the team’s insights with the world. Bird plotted age-of-onset data for 31 familial cases out on a graph. It showed three distinct clusters - families with onset before age 50; families with onset after age 60; and families with variable ages of onset. For example, the Volga German families showed onsets between the ages of 40 - 75. It was not only useful, but actually critical, to group patients by the age they started showing symptoms.
“No genes had been discovered at that time in 1990, but I said, ‘I'll bet that there are at least three genes for Alzheimer's disease’,” says Bird. This statement went against the grain of accepted wisdom in the Alzheimer’s research field at the time.
“[Bird] was the one who opened the door to using age-of-onset [as a key differentiating variable] in our Alzheimer’s research study," says Wijsman. “Everybody else followed eventually, because we were successful, and they weren't.”
“DO-IT-YOURSELF GENOMIC SCIENCE”
The team was hoping that by focusing on subpopulations of early-onset cases, they would have “cleaner” data that would give them a higher chance of success. It was, as Schellenberg remembers, the age of “do-it-yourself” genomic science. “Nowadays, you can view the whole genome sequence online, but back then, the genes were not mapped, and the sequences were not available,” he says. They got a lucky break in that regard thanks to James L. Weber, PhD, a collaborator who was working on the Human Genome Project and amassing DNA markers. A UW laboratory technician named Elaine Loomis went to Weber’s lab to pick out points of interest in the human genome, run the genotypes from the DNA markers, and send those numerical representations of the genetic code to Wijsman for linkage analysis. Then, Schellenberg would use the results to generate a score of the data’s significance for each individual family group. Schellenberg knew that their effort to narrow down the location of a familial Alzheimer’s genetic abnormality to a single chromosome had no real guarantee of success, never mind finding its specific “address”.

'Restriction fragment length polymorphism' technology (left) was used to genotype individuals represented on a medical pedigree (right). Courtesy, Wessex Reg. Genetics Centre
At the end of one long night, Schellenberg processed the one last genotype sent over from Weber’s lab. “And I'm sitting there feeding data into my database and printing it out,” said Schellenberg. “And I started to see big, positive numbers for a linkage to chromosome 14, and I was like, ‘Oh my God!’… That was my ‘Aha’ moment. I had figured that our genetic approach was going to work someday, but the fact that it worked on those early-onset families blew me away.” They were the closest to discovering a gene than they had been since they started in 1986.
In 1992, the UW ADRC published the linkage of a familial Alzheimer’s disease-causing gene to chromosome 14. While this finding did not involve the Volga German family samples, the finding was a big step forward. “I really credit a lot of our success to Tom Bird,” says Schellenberg. Wijsman agrees: “Before the center was even funded, [Bird] was looking at early-onset Alzheimer's families.
His finding of those three clusters of ages of onset, and his suggestion that those represented different biological diseases, focused us on the early-onset families and got us in a place to find the linkage to chromosome 14. I credit those decisions as being partly why we made the progress we did make early on,” says Wijsman. The team’s choice to look at their data in a more selective way helped make success a reality.
AT THE CROSSROADS
At this point, the ADRC linkage study had opened the door to actually identifying the Alzheimer’s genetic risk factor on chromosome 14, kicking up some scientific competition over who could find it first. A group of scientists at Harvard University and Massachusetts General Hospital followed up on the ADRC’s findings and beat them to the identification of the disease gene in several families in 1995. The Harvard team named the gene “presenilin-1” (PSEN1) because the newly identified mutation in this gene nearly always led to dementia in people’s 40s, at much younger ages than for the other genetic variants of Alzheimer’s disease.
Despite the disappointment of not finding the PSEN1 mutation first, the ADRC’s linkage of the mutation to DNA on chromosome 14 allowed for the Alzheimer’s research field to come to terms with larger, looming challenges. “Our finding really disturbed people because it showed that the genetics weren’t going to be simple – that there would be more than one gene for familial Alzheimer’s,” says Bird. “The field wanted it to be simple, and everyone was disappointed that it wasn’t going to be.” Steering in a direction that would characterize the next two decades of ADRC research, the early molecular genetics team embraced the emerging complexities of Alzheimer’s disease.
In a landmark paper published in 1989, Evidence for Etiologic Heterogeneity in Alzheimer's Disease, the team had suggested that there are multiple different genetic mutations behind familial Alzheimer’s disease. They posited that while some families carry a mutation on chromosome 21, there would be other familial Alzheimer’s mutations to discover still. The paper went beyond genetic mutations to introduce the influential idea of ‘heterogeneity’ in Alzheimer’s disease, or the concept that several different genetic and environmental factors, such as toxins, trauma, or viruses, may act separately, or in combination, to result in a cognitive syndrome linked to the pathology of amyloid plaques and tau tangles. Genetic factors may act differently in the context of an individual's own biology and health issues, explaining the variability in the symptoms and ages of onset in familial cases, as well as the sporadic, later-onset forms of Alzheimer’s disease.

For example, the ADRC team noticed that the Volga German families in their study showed wider variability in the ages of onset than other family groups. Some people developed symptoms at the age of 40 while others did not until age 72. The types of cognitive symptoms and duration of illness also differed from person to person.
In the landmark 1989 paper, the authors suggested that the distinct manifestation of familial Alzheimer’s involves factors beyond the gene —even in cases in which a genetic factor was so strong that virtually all people with that mutation would someday develop dementia. This phenomenon of varying levels of resilience energized the idea that it is just as important to study the factors that counter disease pathology in the brain as it is to study the factors that cause it.
ADRC scientists of this early era honed in on the idea that each person has a different combination of biological and environmental factors that influence risk and resilience to neurodegenerative diseases and dementia. This concept of 'biological heterogeneity,' from a genetic perspective, was the central theme of the original ADRC – and the researchers soon discovered that the genetics of Alzheimer’s disease would be more complicated than anyone expected. Now, in 2020, the newly renewed ADRC continues a thematic focus on biological heterogeneity, using new tools such as pluripotent stem cells. These in-vivo models allow researchers to probe the mechanisms of genetic variations in the context of an individual patient’s biology and identify therapeutic drug targets.
ADRC researchers now study genes whose variants may contribute some level of risk or resilience to Alzheimer’s: UBC, SORL1, MSUT2, and APOE, as well as newly identified risk factors of frontotemporal degeneration and ALS. In this light, the story of the early ADRC embodies the evolution of the field - from a view of Alzheimer’s disease as a single gene phenomenon, to a condition that varies across individuals involving many different underlying factors that converge at cognitive trouble, and ultimately, will require different treatments.
COLLABORATION LEADS TO DISCOVERY
In 1994, the team was still looking for the genetic cause of Alzheimer’s disease in the affected Volga German families. Schellenberg and Wijsman had ruled out the amyloid gene on chromosome 21 and the presenilin gene on chromosome 14. The data suggested the Volga German families carried a third, as-of-yet-undiscovered gene for Alzheimer’s disease—a gene that had been flying under the researchers’ radar. Eager to bring new energy to the Volga German family mystery, Schellenberg asked Ephrat Levy-Lahad, an up-and-coming post-doctoral researcher in the medical genetics program, to head the project. Under Schellenberg’s guidance, Levy-Lahad prepared and sent the genotypes to Wijsman, who started digging through the data for a correlation between DNA and disease. They started making progress immediately.
“I remember Ellen [Wijsman] came to me with the news,” says Bird. “She said, ‘We’ve got it – a linkage in these families. [Their disease gene] is linking to the long arm of chromosome 1. It’s pointing to a new gene for Alzheimer’s disease. We’ve got to report this.” They wrote up a paper and Science started reviewing it.
After seven years of trying, genetic linkage studies on these Volga German families had finally led the group to an area on chromosome l, but they had still not identified the gene itself. “We couldn't look it up—nobody knew what genes were in that particular region,” says Schellenberg.

Levy-Lahad refers to the technology that they used at the time as “prehistoric”. “Genetics has undergone a total revolution since then. Today we have the whole human genome, we know where all the genes are, and we can sequence them in a week,” says Levy-Lahad. “Telling my graduate students today about the technologies we used to sequence genes back then would almost be like describing how we used to have to mine our own salt or gather our own coals.”
Mapping out the different genes within that region on chromosome 1, at the time, required yeast artificial chromosomes, called “YAC” clones, which are human-engineered DNA molecules used to clone DNA sequences in yeast cells. This technology would allow the researchers to run many experiments on their DNA sequence of interest. Fortunately, Maynard Olson, PhD, now Professor of Genome Sciences and of Medicine, had arrived at the UW in 1992 and created an impressive YAC clone library, now at the ADRC’s disposal.
The team also had help organizing the YACs that they would need to cover the linked region on chromosome 1, thanks to Chang-En Yu, PhD, a researcher who still works as Research Associate Professor in the Division of Gerontology & Geriatric Medicine at the VA.
Levy-Lahad got to work. “We had to take these YACs and start sequencing them to try and figure out which genes were there,” she says. “At first, we were going through them, one by one, just like we had been doing for the chromosome 14 gene.” It took sixty YACs to cover the candidate region on chromosome 1; finding the genetic mutation would be akin to searching for a needle in a haystack.

Ephrat Levy-Lahad, MD
It is at this moment that external collaboration became a rung on the ladder of scientific discovery, with a call from Wilma Wasco, PhD and Rudolph Tanzi, PhD, who had both worked with the Harvard team that followed the ADRC's lead to identify the mutation in the presenilin-1 gene on chromosome 14 in their group of early-onset families.
In spite of the previous competition, they collaborated with the ADRC team and gave them a clue that would prove critical in their probe into the genetics of the Volga German families: Wasco and Tanzi had recently found a genetic fragment with a sequence highly similar to the presenilin-1 gene located on chromosome 14. The fragment was a “homolog,” or a genetic cousin of the chomosome 14 gene. The similarity between the two gene sequences suggested that the proteins they encode may have similar functions that could be important to Alzheimer's biology.
The implication of Tanzi’s message was that, perhaps, searching for this homolog in the Volga German family DNA may lead the ADRC team to the mutation site. It was far-fetched. The ADRC team was skeptical.
“The significance of the homolog was just that there was something specific to look for,” says Levy-Lahad. "If you would have asked Jerry and I, we would have said that the chances looked really slim.” She searched the YACs in the chromosome 1 region for the homolog gene, and, remarkably, she found it. The homolog was right there, hiding away in the DNA region that the team had pinpointed as the likely site of the Alzheimer's variant in the Volga German family members."That was like a sledgehammer to the forehead," Schellenberg told medical reporters at the time. "It went from being a ho-hum project to... 'this is the gene'."
Schellenberg calls the subsequent search to find the gene in the Volga German families “a slam dunk.” He still remembers the moment of discovery, when Levy-Lahad and Loomis were looking at the genetic sequencing data that came out on giant X-ray films, which measured about 14 by 18 inches. “I saw them holding this thing and going into the lab, laughing and giggling and smiling,” he says. “They wouldn't tell me, but I knew they had the gene.”
“There's so much of an element of luck in these things,” says Levy-Lahad. “I was working 18-hour days to find this gene, and it could have been a project of a few years if there wasn't that homolog.”
The team finished the job within a few days, with help from David Galas, PhD and colleagues at Darwin Molecular, a biotech firm in Bothell, Washington. They were able to sequence the gene from the Volga German family members and, critically, confirm that the mutation was only present in the gene in affected family members and absent in unaffected members. “The genetics all fit together so nicely,” says Schellenberg. “It was incredibly rapid.”
A small genetic change had proven responsible for the Volga German families’ history of Alzheimer’s disease: a single mutation, N141I, within the gene that was named “presenilin-2” for its similar biological structure to the presenilin-1 gene. The ADRC researchers described the discovery in the August 18, 1995 issue of the journal Science, and received coverage in Science Magazine.

Dimensions cover story of Fall 1995
The team toasted their achievement. Their decade-long effort had paid off in the discovery of a new Alzheimer’s gene mutation, fulfilling the aim set out in the original 1985 ADRC application to identify important genetic factors. “The VA didn’t allow alcohol into the building,” said Levy-Lahad, “So we took our bottle of champagne and celebrated in the parking lot.” The popping of the cork didn’t just mark one scientific achievement. It ushered in an era of even more fruitful and expansive research in Alzheimer’s disease genetics.
A year later in 1996, the team published another genetic study on presenilin-1 and presenilin-2 that is now the most highly cited paper in ADRC history. In 1998, the genetics group identified a mutation in the Tau gene, and published the first report of a genetic cause of the neurodegenerative disease frontotemporal degeneration. Returning to the study of presenilin-1 in 1999, ADRC researchers created the first mouse model of dysfunction in this gene, opening up a new avenue for investigating the biology of the Alzheimer’s-causing gene variant in a living organism.
In the current day, presenilin work is as productive as ever and directly proceeds from the studies of the 80s and 90s. Bird’s protégé, Suman Jayadev, MD, Associate Professor of Neurology and ADRC Clinical Core Lead, and her team recently uncovered evidence that age-related changes in the way presenilin-2 generates transcripts (the instructions for making proteins) may be involved in later onset Alzheimer’s risk in the general population. Now, an ADRC Development Project is deploying the newest genetic technology to expand the understanding of presenilin-1 and 2 variants in different forms of both familial and sporadic late-onset Alzheimer's disease, which may reveal new functions for these genetic factors. The researchers see this project as a possible first step on the path to a new treatment involving the preservation of genetic function.








